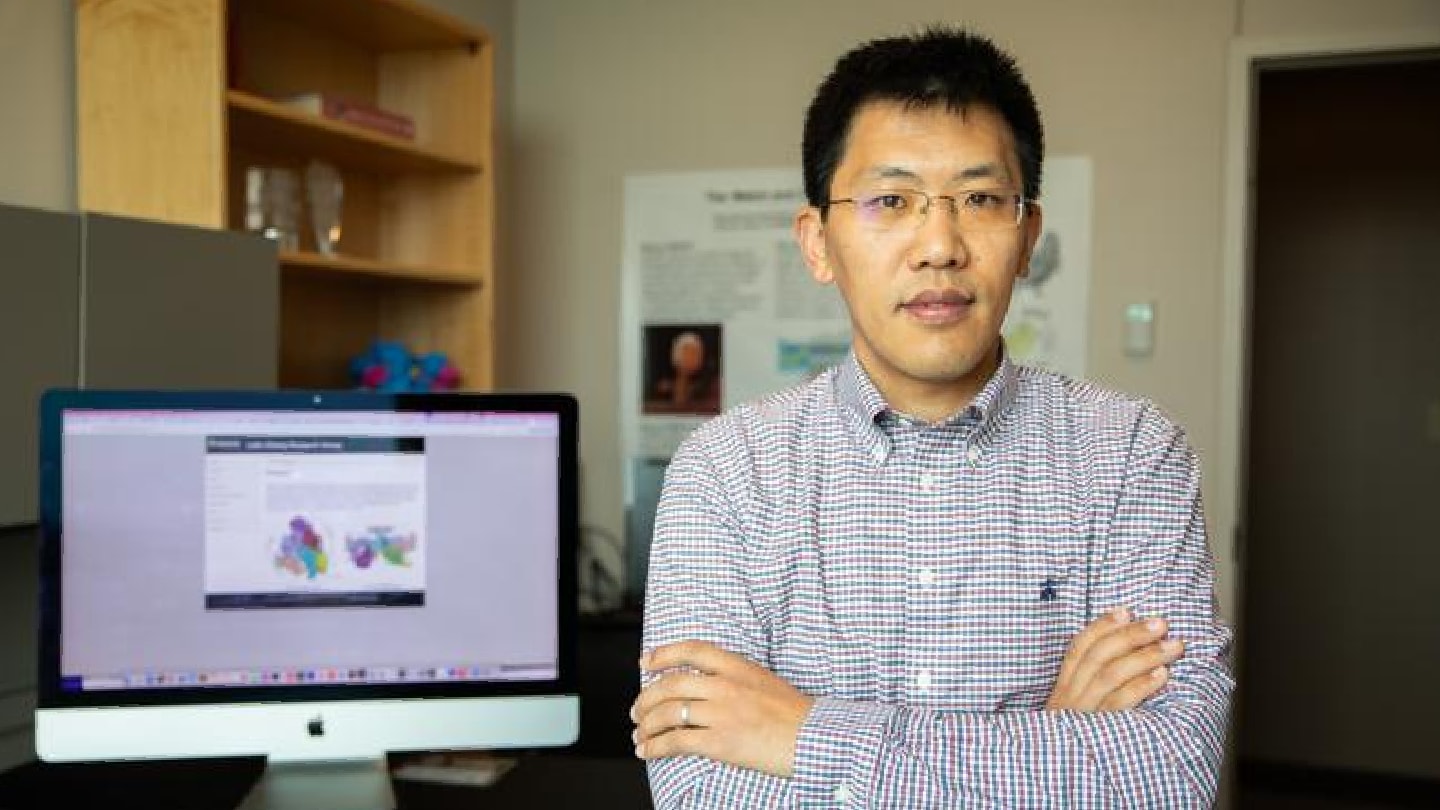
Homegrown CAR T Therapy Clears FDA Hurdle for AML Trial
University of Colorado Anschutz launches first U.S.-cleared, campus-built CAR T program for adults with relapsed or refractory acute myeloid leukemia
The University of Colorado Anschutz’s Gates Institute has won FDA clearance to begin clinical testing of a CAR T-cell therapy developed entirely on campus – the first U.S. program of its kind to reach that stage. The treatment targets CD64, a protein found on aggressive acute myeloid leukemia (AML) cells that can persist after modern therapies, and will now move into a first-in-human Phase 1 trial.
The study, expected to begin this summer, will enroll adults with relapsed or refractory AML at UCHealth University of Colorado Hospital. Investigators will assess the therapy’s safety, tolerability, and optimal dose. A pediatric trial is also planned for later in 2026, potentially expanding the approach to children and adolescents with hard-to-treat disease.
The program highlights the growing ambition of academic cell therapy centers to take treatments from discovery to manufacturing and early clinical testing under one roof. The CAR T product for the trial will be made at the Gates Biomanufacturing Facility, creating a direct pipeline from lab bench to bedside for a patient group still facing limited options and poor outcomes.
CAR T Therapy Targets Tumor Ecosystems
uPAR-directed CAR T cells dismantle fibrotic, immunosuppressive niches and show broad efficacy across solid tumors
A new study suggests that one of the biggest barriers to CAR T therapy in solid tumors – an inhospitable tumor microenvironment – may be overcome by targeting not just cancer cells, but the ecosystem that sustains them. Researchers identify the receptor uPAR as a unifying marker of aggressive tumors and their surrounding fibrotic, immunosuppressive niches, opening the door to a more comprehensive immunotherapy strategy.
Across more than 10,000 patient samples, uPAR was widely expressed in solid tumors – particularly those with TP53 and RAS pathway mutations – and associated with poor prognosis. Crucially, uPAR was found not only on tumor cells but also on neighboring fibroblasts and immune cells in a senescence-like state, forming a protective niche that limits immune attack. CAR T cells engineered to target uPAR eliminated both compartments, achieving durable tumor regressions across multiple cancer models, including metastatic disease.
In mouse and humanized models, a single infusion of uPAR CAR T cells cleared tumors with minimal long-term toxicity, despite targeting some normal myeloid cells. The approach also showed synergy with chemotherapy: senescence-inducing treatments increased uPAR expression, boosting CAR T cell efficacy and leading to long-term survival in most treated animals.
By focusing on a shared tumor state rather than a lineage-specific antigen, the strategy sidesteps the heterogeneity that has hampered CAR T therapies in solid cancers. The findings position uPAR as a rare dual-purpose target – one that enables simultaneous tumor killing and microenvironment remodeling – and suggest broader applications beyond oncology, including fibrotic and age-related diseases.
Base Editor Redesign Slashes Off-Target Mutations Without Sacrificing Power
A single amino acid tweak transforms a high-efficiency gene editor into a safer therapeutic candidate
A widely used gene-editing tool may be far less safe than assumed – but a simple redesign could fix that. Researchers report that ABE8e, one of the most efficient adenine base editors, introduces genome-wide off-target mutations at rates up to 30-fold higher than natural background levels in mouse embryos. These unintended edits occur independently of guide RNA, raising concerns for therapeutic use.
To address this, the team systematically re-engineered the enzyme’s deaminase domain. Their standout variant, ABE8eY149V, retains the high editing efficiency of ABE8e while dramatically reducing off-target effects to near-background levels – both in DNA and RNA. The improvement stems from a single amino acid substitution that narrows the editor’s activity window and limits nonspecific interactions.
The upgraded editor also proved versatile, working with multiple CRISPR systems to expand target range. In human cells, it corrected disease-relevant mutations with greater precision than existing variants. In a mouse model of hereditary tyrosinemia type I, it restored liver function and prevented death after in vivo delivery.
Base Editing Reverses Deadly Epilepsy Mutation in Mice
A single-shot gene-editing therapy restores neuronal function, eliminates seizures, and prevents early death in a severe SCN8A epilepsy model
A CRISPR-derived base editing therapy has rescued seizures and dramatically improved survival in a mouse model of SCN8A developmental and epileptic encephalopathy (DEE). The approach directly corrects a recurrent gain-of-function mutation (R1872W) in the Nav1.6 sodium channel gene – one of the most severe and treatment-resistant genetic epilepsies.
Delivered via dual AAV vectors shortly after birth, the adenine base editor converted the disease-causing mutation back to its healthy sequence in a subset of neurons. Despite editing only ~32% of mutant transcripts, the treatment eliminated seizures entirely in most mice and significantly reduced seizure frequency and severity in the rest. Survival improved markedly, with ~87% of treated animals living beyond typical lethal timepoints, while untreated mice died early from seizure-induced events.
Electrophysiology revealed that the therapy normalized neuronal hyperexcitability and suppressed the pathological persistent sodium current (INaP), a key driver of seizure activity. Behavioral deficits – including reduced mobility and anxiety-like traits – were also improved. Importantly, transcriptome and whole-genome sequencing showed minimal off-target editing (≤1%), underscoring the precision of the approach.
RNA-Guided Gene Activation Found in Nature
Researchers uncover a bacterial system in which nuclease-dead Cas12f and a sigma factor team up to switch genes on without conventional promoter sequences.
CRISPR proteins are best known for cutting DNA or, in engineered form, turning genes off. Now, researchers have identified a natural system that does the opposite. Writing in Nature, Florian T. Hoffmann and colleagues describe nuclease-dead Cas12f homologues that partner with an extracytoplasmic function sigma factor, σE, to drive RNA-guided transcription. Rather than relying on standard promoter motifs, the complex uses guide RNA to bring σE and RNA polymerase to DNA targets marked by a minimal 5′-G target-adjacent motif, triggering transcription at a fixed distance from the Cas12f-bound site.
The team showed that dCas12f binds guide RNAs and target DNA, recruits σE in a guide-dependent fashion, and can activate gene expression in E. coli when paired with its cognate RNA polymerase. Strikingly, transcription start sites were positioned almost entirely by geometry: they consistently appeared about 46 base pairs downstream of the target-adjacent motif. By redesigning the guide RNA, the researchers could shift transcription start sites across the genome and even drive transcription in either direction, including from loci lacking recognizable promoter elements.
Bioinformatic and genomic analyses suggest these systems naturally regulate transporter operons and signaling pathways, especially in Bacteroidetes, where sigma factor networks are already unusually elaborate.